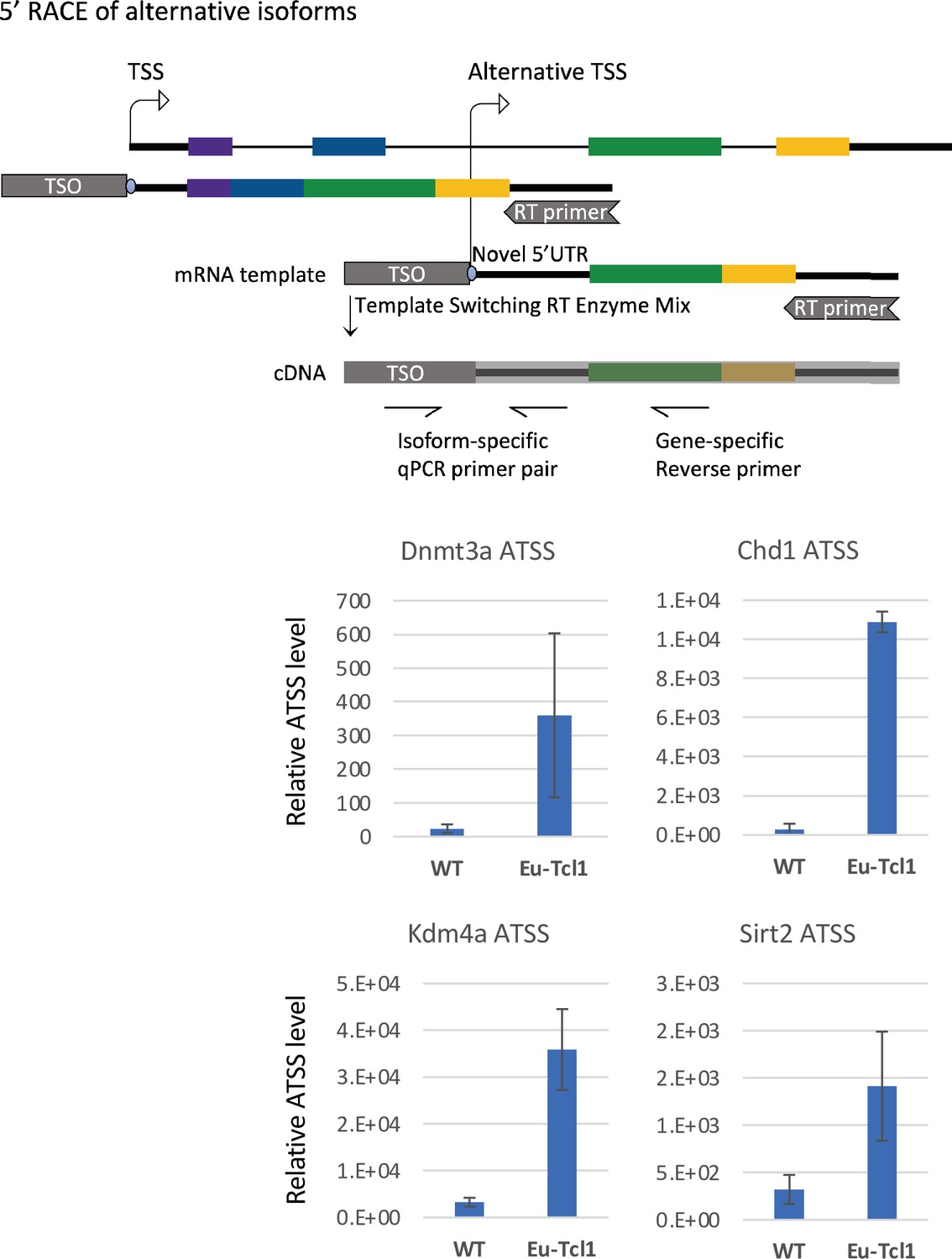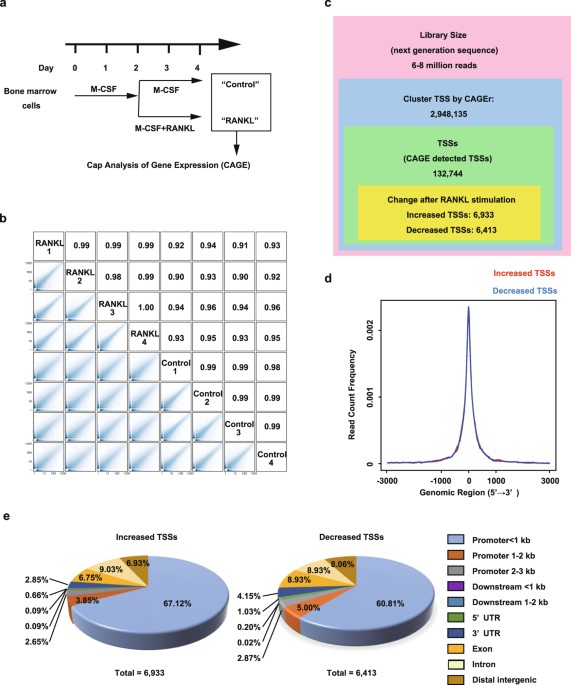
Roles of Enhancer RNAs in RANKL-induced Osteoclast Differentiation Identified by Genome-wide Cap-analysis of Gene Expression using CRISPR/Cas9 | Scientific Reports
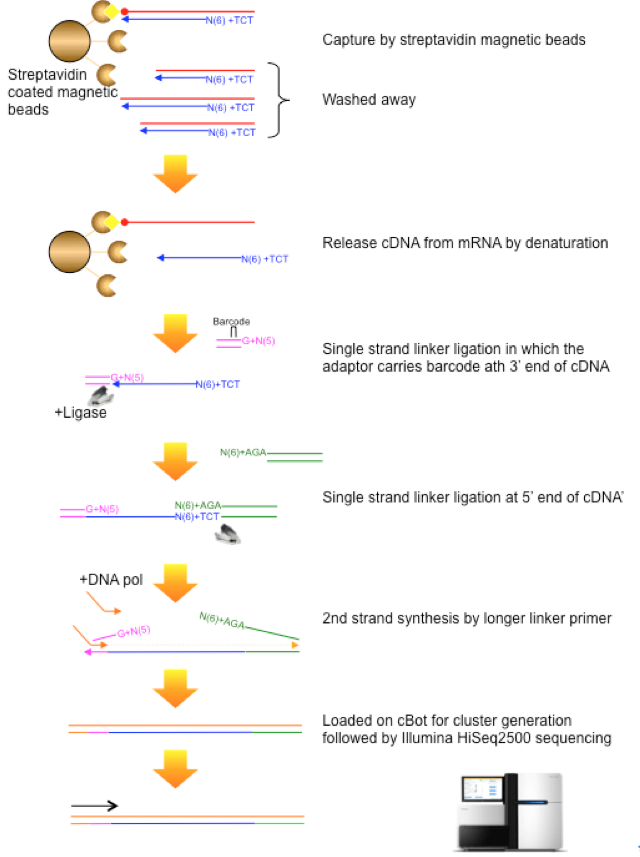
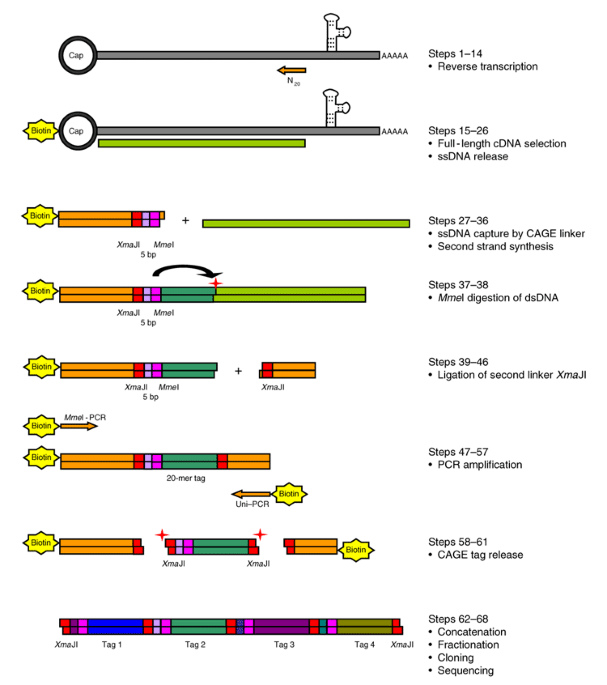
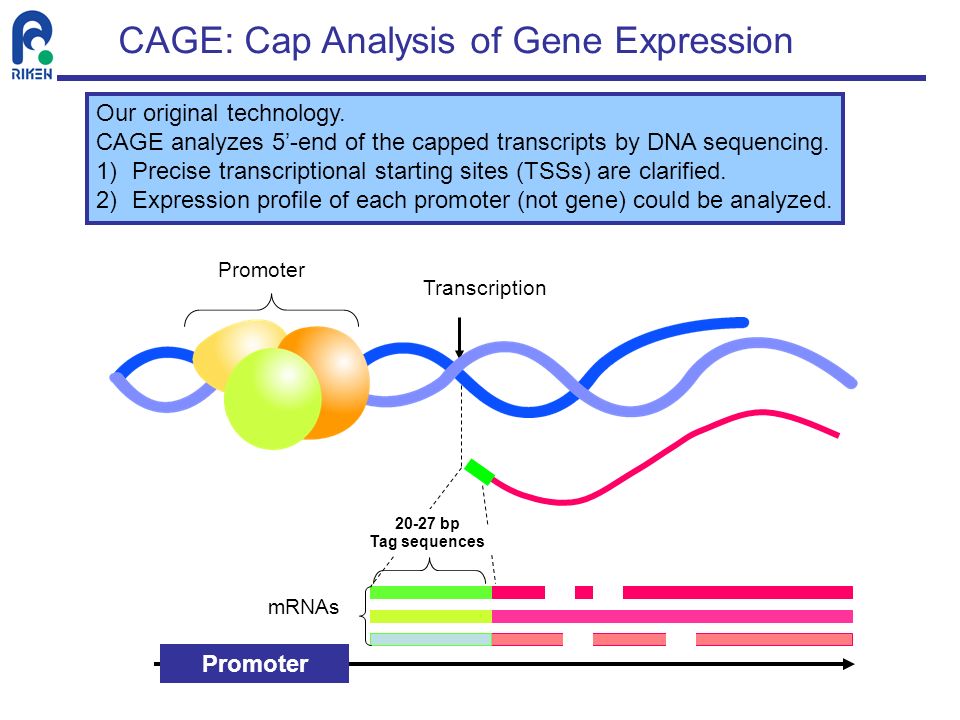
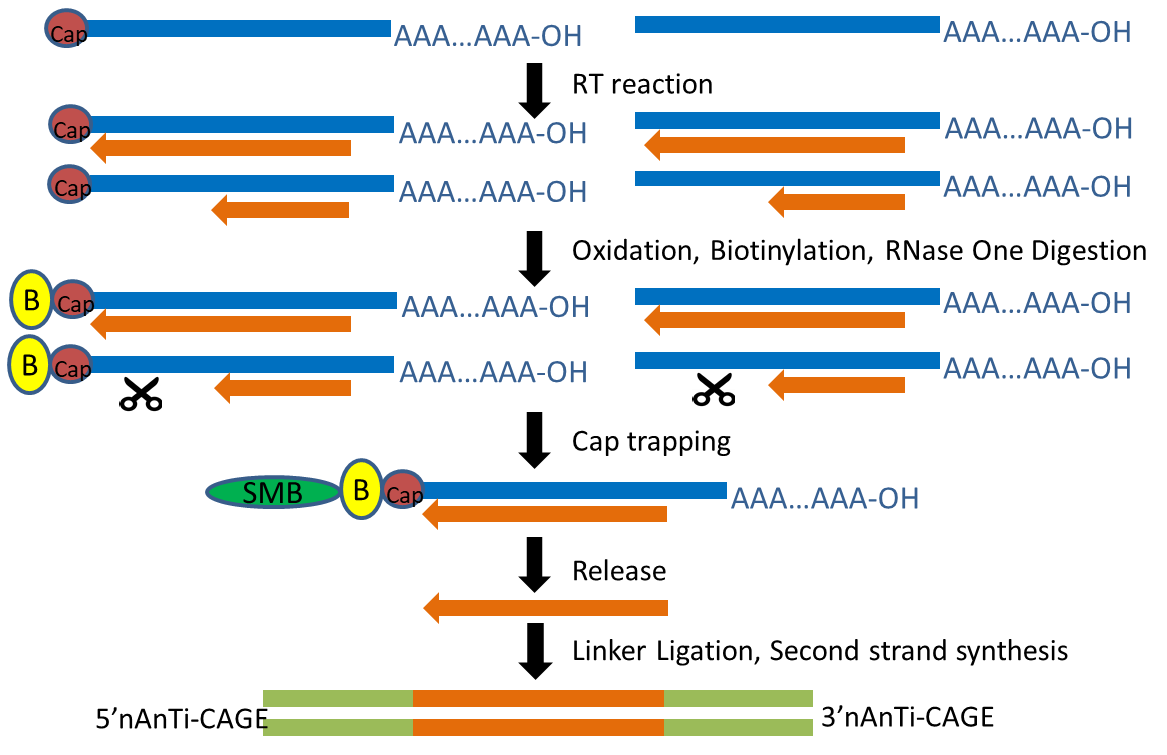
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター
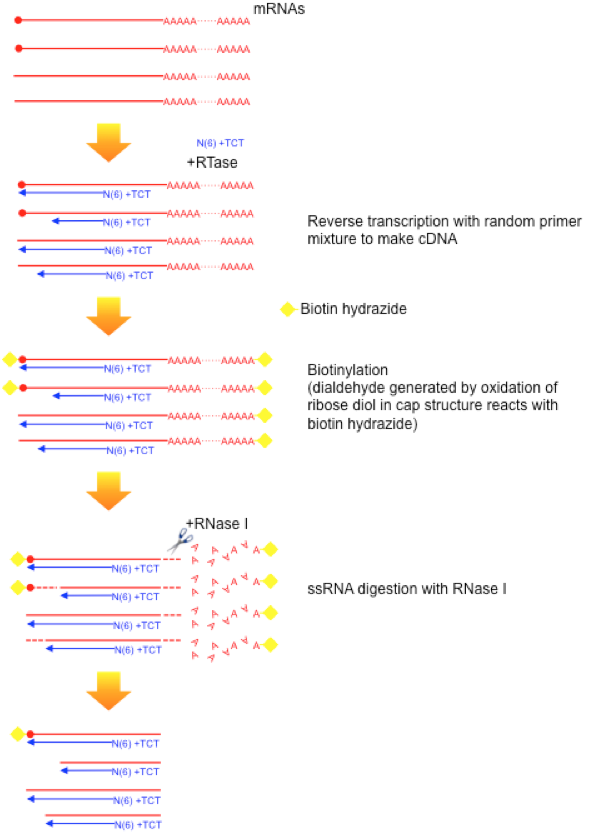
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター

Cap-analysis of gene expression (CAGE) analyses of transcripts during... | Download Scientific Diagram
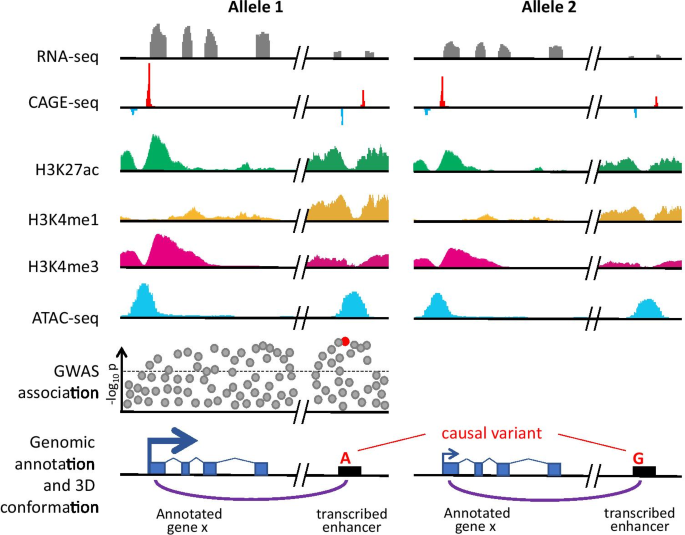
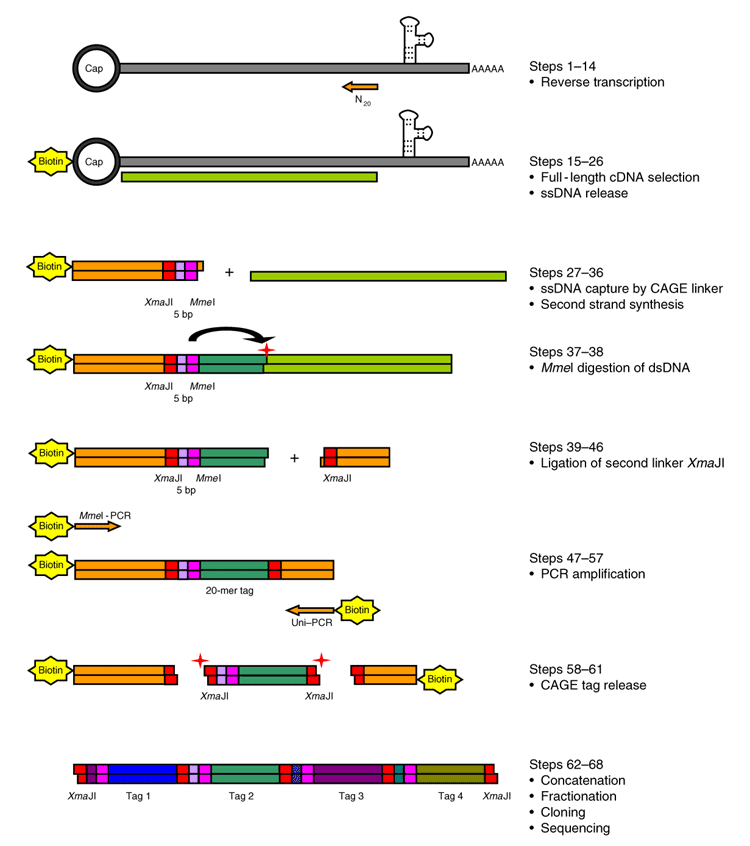
Cap analysis of gene expression (CAGE) and noncoding regulatory elements | Seminars in Immunopathology

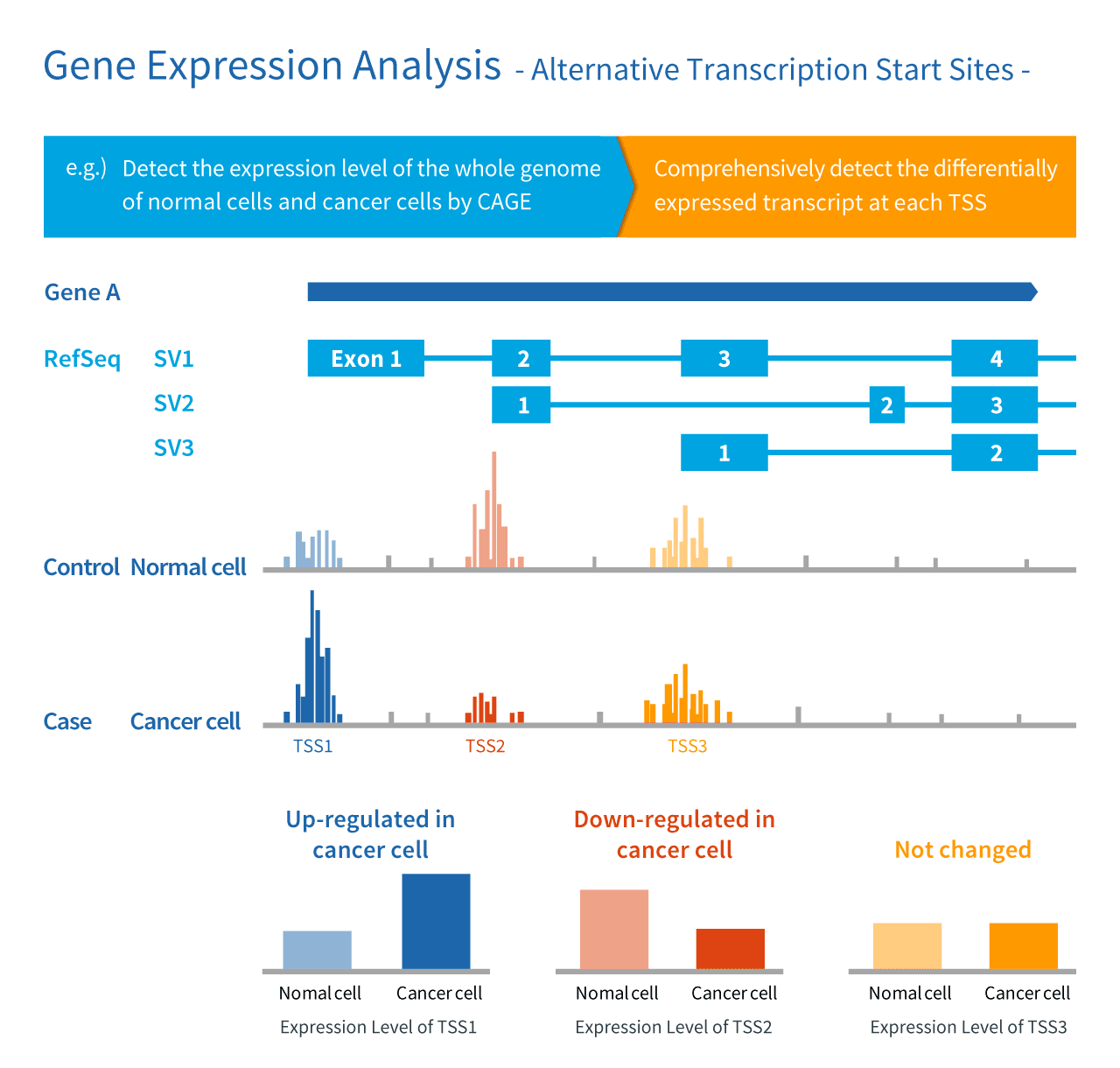
Gene expression profile of HN by RNA-Seq CAGE (Cap Analysis of Gene... | Download Scientific Diagram
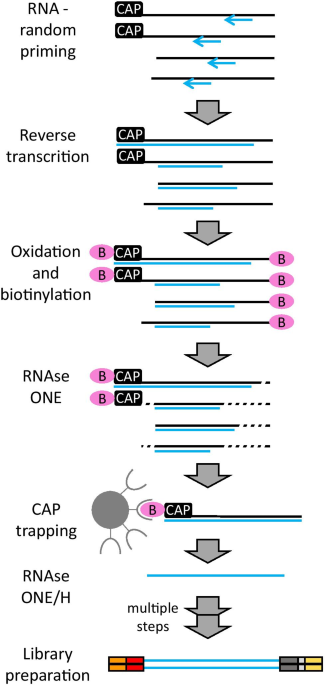
Cap analysis of gene expression (CAGE) and noncoding regulatory elements | Seminars in Immunopathology